| structure name | CRYSTAL STRUCTURE OF THE INTEIN HOMING ENDONUCLEASE PI-SCEI BOUND TO ITS SUBSTRATE DNA (CA2+ FREE) (ENDONUCLEASE PI-SCEI) |
| reference | Quiocho et al. |
| source | Saccharomyces cerevisiae |
| experiment | X-ray (resolution=3.20, R-factor=?) |
| structural superfamily | Hedgehog/intein Hint domain;Homing endonucleases; |
| sequence family | Hom_endassociated Hint; Homing endonuclease; |
| redundant complexes |  1lws_A 1lws_A
|
| links to other resources | NAKB  PDIdb DNAproDB PDIdb DNAproDB |
| protein sequence | |
| interface signature | QRLRIKYEESSSHKRYKNEKDNTHY |
Estimated binding specificities ?
| readout + contact | 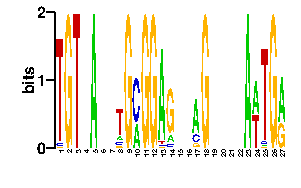 |
A | 0 0 84 63 0 24 21 24 24 0 7 24 24 5 1 0 0 0 0 84 24 24 0 24 96 0 92 C | 23 96 8 1 0 24 59 24 24 96 1 24 24 78 4 96 96 0 96 3 24 24 0 24 0 96 0 G | 1 0 1 0 0 24 13 24 24 0 17 24 24 12 1 0 0 60 0 6 24 24 0 24 0 0 3 T | 72 0 3 32 96 24 3 24 24 0 71 24 24 1 90 0 0 36 0 3 24 24 96 24 0 0 1scan! |
Dendrogram of similar interfaces ?
matrix formatQGRALARGITKAYTETECSTSTSTHTKGRG------YA----KG------NTECKCDGNCTTHG--YT L +---------------------1lwt_A +------2 ------------------------------QGEC--------QC------------------------ L ! ! +-------------3c0w_A --3 +-------1 --------------------------------TGQG--RGRGWC--------------------VC-- L ! +-------------2vs7_A ! --------------------------------------------KGCGFT------------------ L +----------------------------2bop_A |
home
updated Sat Jan 10 04:47:42 2026
